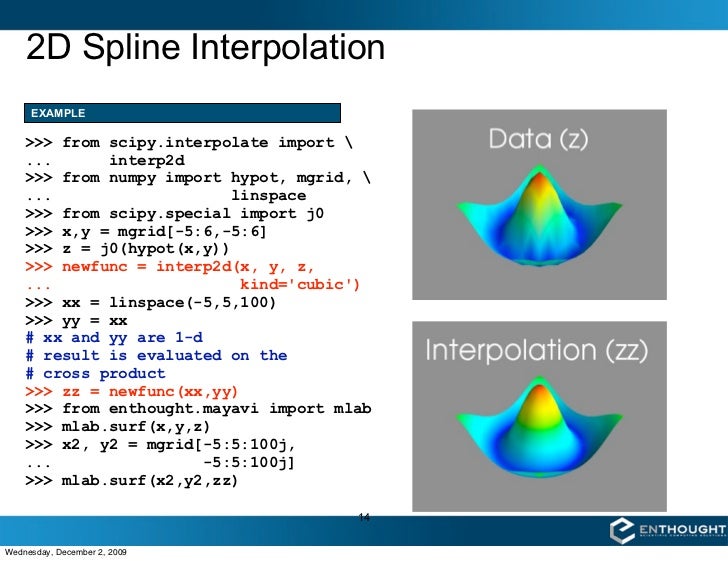
 

Thank you for the hints in your question. version = 0.14.0, it happily works on some 1600x1600 model output. I get the same error as you do with scipy. I think older versions assume irregular grids and don't take advantage of the regular grids. astype(band_array.dtype) to make the output data type the same as the input array. 
Note that the result will be returned with an apparent higher precision than the source data, since it is up-classed to NumPy's dtype('float64') data type. # Use the differences to weigh the four raster values # Calculate differences from point to bounding raster midpoints # Test array bounds to ensure point is within raster midpoints # Special case where point is on upper bounds # Calculate raster lower bound indices from point I've translated the formula below (from Wikipedia) into Python-speak to yield the following algorithm, which appears to work. but I much prefer bilinear interpolation methods) (Note: the nearest neighbor interpolation algorithm is easy cake: from numpy import argmin, NANĭef nearest_neighbor(px, py, no_data=NAN): Has anyone come across a good bilinear interpolation algorithm, preferably in Python, possibly tailored with NumPy? Any hints or advice? I don't see why I should have a memory error, since my raster is 317×301, and the bilinear algorithm should not be difficult. I'll admit, I have limited confidence in this SciPy function, as the bounds_error or fill_value parameters don't work as documented. Self.tck = fitpack.bisplrep(self.x, self.y, self.z, kx=kx, ky=ky, s=0.)įile "C:\Python25\Lib\site-packages\scipy\interpolate\fitpack.py", line 873, in bisplrep However I get a memory error on my 32-bit Windows system with a 317×301 raster: Traceback (most recent call last):įile "C:\Python25\Lib\site-packages\scipy\interpolate\interpolate.py", line 125, in _init_ Up to now, I've tried SciPy's interp2d function: from scipy import interpolateīilinterp = interpolate.interp2d(ax, ay, band_array, kind='linear') Nx, ny = source.RasterXSize, source.RasterYSizeīand_array = source.GetRasterBand(1).ReadAsArray()Īx = array( ix*gt gt/2.0 for ix in range(nx)])Īy = array( iy*gt gt/2.0 for iy in range(ny)]) Here is where I'm at: from osgeo import gdal Z = np.I have a raster that I'd like to do some point interpolations with. # I'm fairly sure there's a more efficient way of doing this. But if it is, you can avoid having to interpolate: def method_3(): 
In general, with sampled data, this will not be true. For every possible unique (x,y) pair, there is a corresponding (x,y,z) in your data.įrom this, it follows that the number of (x,y,z) pairs must be equal to the square of the number of unique x points (where the number of unique x positions equals the number of unique y positions).The number of different x sample positions equals the number of different y sample positions.There's a third option, depending on how your (x,y,z) is set up. It may have one or more negative-space holes which are also bounded by linear rings. A polygon is a two-dimensional feature and has a non-zero area. Method 3: No Interpolation (constraints on sampled data) A geometry type representing an area that is enclosed by a linear ring. This method produces the following graphs: Z = iddata((mat, mat), mat, (X,Y), method='nearest') # Depending on your "error", you may be able to use other methods # Interpolate (x,y,z) points over a normal (x,y) grid # This works if you have (x,y,z) tuples that you're *not* generating, and (x,y) points Method 2: Interpolating given z points over a regular grid # Method 2: Here, the returned matrix looks like: mat = , Y_err = (np.random.rand(*y.shape) - 0.5) * xy_max_error X_err = (np.random.rand(*x.shape) - 0.5) * xy_max_error GEOS, a port of the Java Topology Suite (JTS), is the geometry engine of the PostGIS spatial extension for the PostgreSQL RDBMS. # Half of this will be in the direction, half will be in the - dir. Shapely is a Python package for set-theoretic analysis and manipulation of planar features using functions from the well known and widely deployed GEOS library. # First we generate the (x,y,z) tuples to imitate "real" data First, I define a function that generates fake data: def gen_fake_data(): If you don't have that ability, and are given a fixed (x,y,z). This is relatively easy, since you can generate z at whatever points you want. # This works if you are generating z, given (x,y) # dim_? is the granularity in that direction If you just have just a list of ( x, y, z) tuples, it's harder (see method_2() below, and maybe method_3()).Ĭonstants # min_? is minimum bound, max_? is maximum bound, you know the formula for it) it's very easy (see method_1() below). Depending on whether you're generating z or not, you have at least two different options. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
